Application for exploiting unstructured data from Computerized Patient Records (CPR) based on medical reports and prescriptions, in collaboration with the Public Interest Group SIB. The system leverages OCR and natural language processing to support clinical decision-making.
1. System Architecture
The project relies on a modular architecture integrating several technical areas:
- Patient record indexing: extraction and structuring of medical information via an LLM model
- Patient triage: classification based on criticality, using an XGBoost model and decision trees
- Length of stay prediction: estimation of hospitalization time based on medical history and admission data
- Interface and API: creation of an API enabling interaction with modules through an ergonomic user interface
2. Project Management
The project followed a rigorous methodology with detailed planning and structured documents. The documents that served as the basis for this planning are the specifications we drafted in agreement with the client and the following functional requirements diagram:
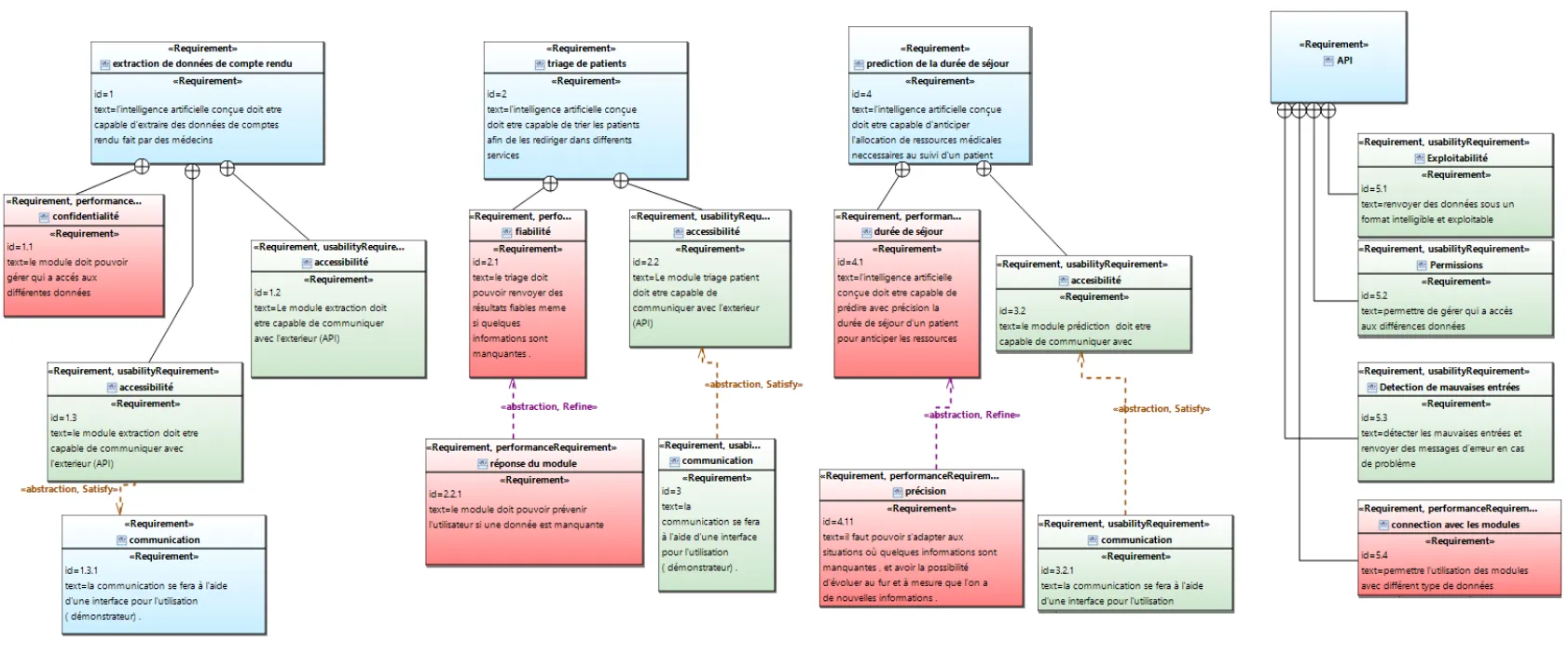
Figure 1 - Functional Requirements Diagram
a) Team Organization
The team was divided into specialized units with clear responsibilities:
| Unit | Responsibilities |
|---|---|
| Indexing | Extraction and structuring of medical data |
| Triage | Development of the classification model |
| Length of Stay Prediction | Implementation of the prediction model |
| Interface & API | Development and integration of the API |
b) Gantt Chart
The project was structured in several phases:
| Phase | Objectives | Duration |
|---|---|---|
| S5 | Training and objective definition | 4 weeks |
| S6 | Development of functional modules | 8 weeks |
| S7 | Validation and improvement of features | 6 weeks |
This allowed us to create a Gantt chart for the entire project duration, which was modified as progress was made and to which we adhered:
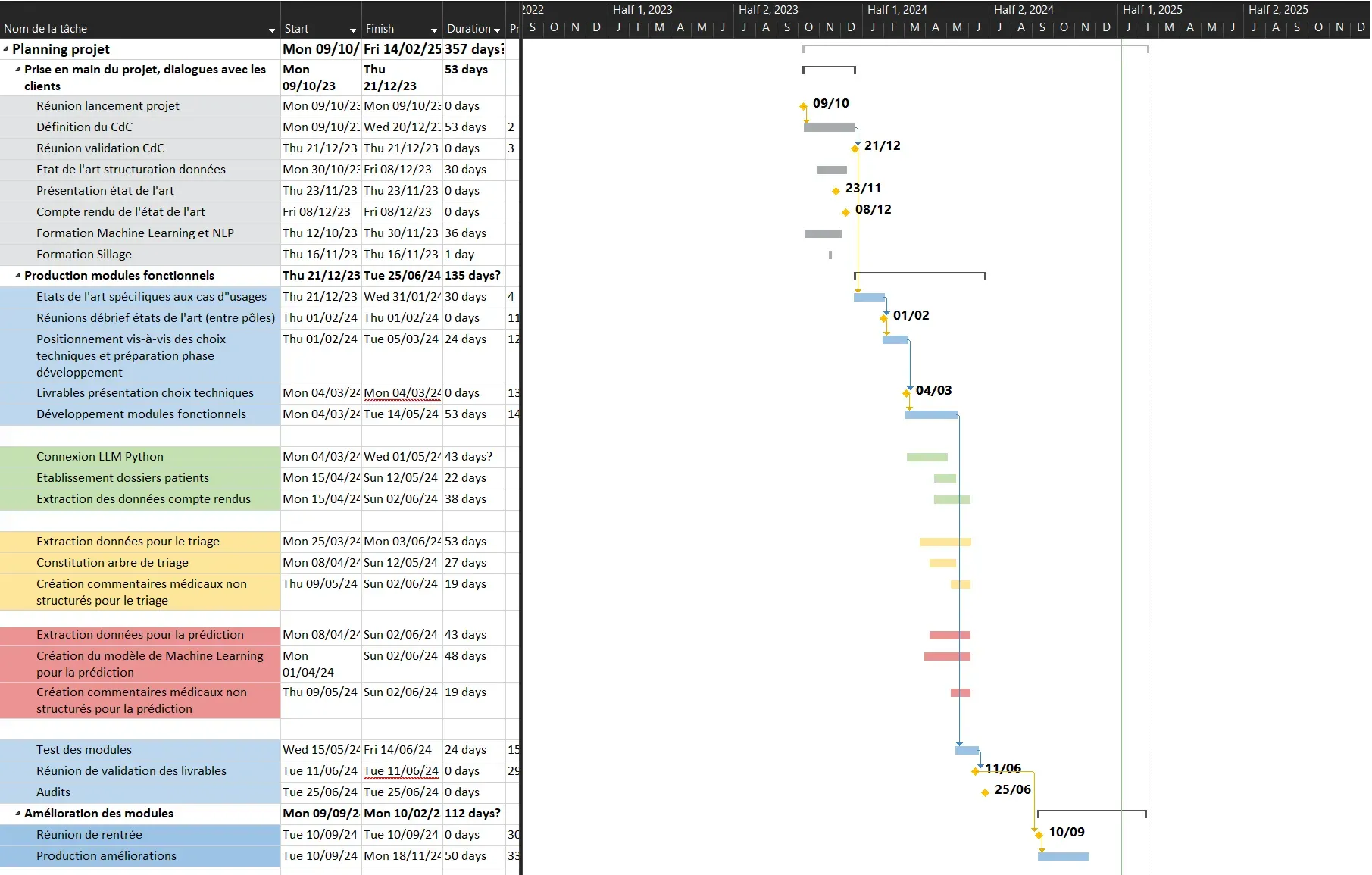
Figure 2 - Gantt Chart
c) Risk Management
A SWOT analysis was conducted to identify project risks and opportunities:
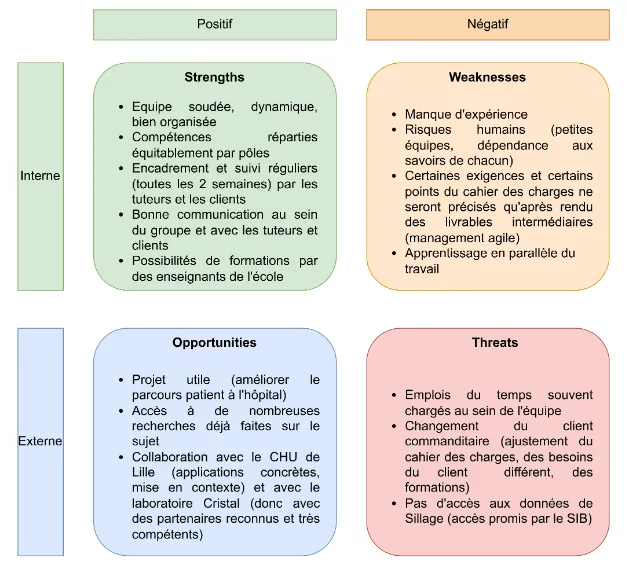
Figure 3 - SWOT Matrix
Additionally, we also added a risk management plan that was also modified during our work:
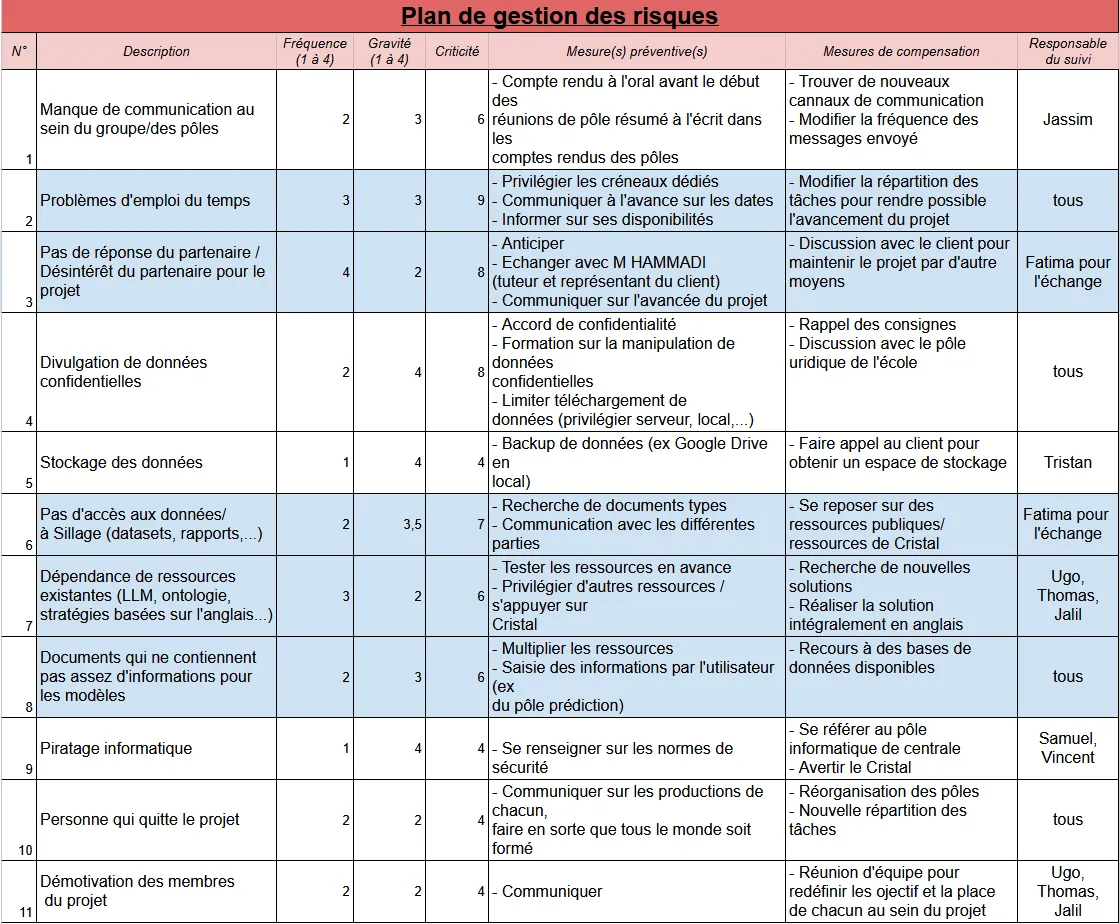
Figure 4 - Risk Management Plan
d) Work Breakdown Structure
This task breakdown was divided into 2 major parts, just like our project: the first includes familiarization with the project and deliverable creation, while the second focuses on their improvement and API usage.

Figure 5 - Work Breakdown Structure
e) RACI Matrix
After all this preparation work and to be able to start the technical part of this project, we drafted a RACI matrix that defines the role of each stakeholder in this project:
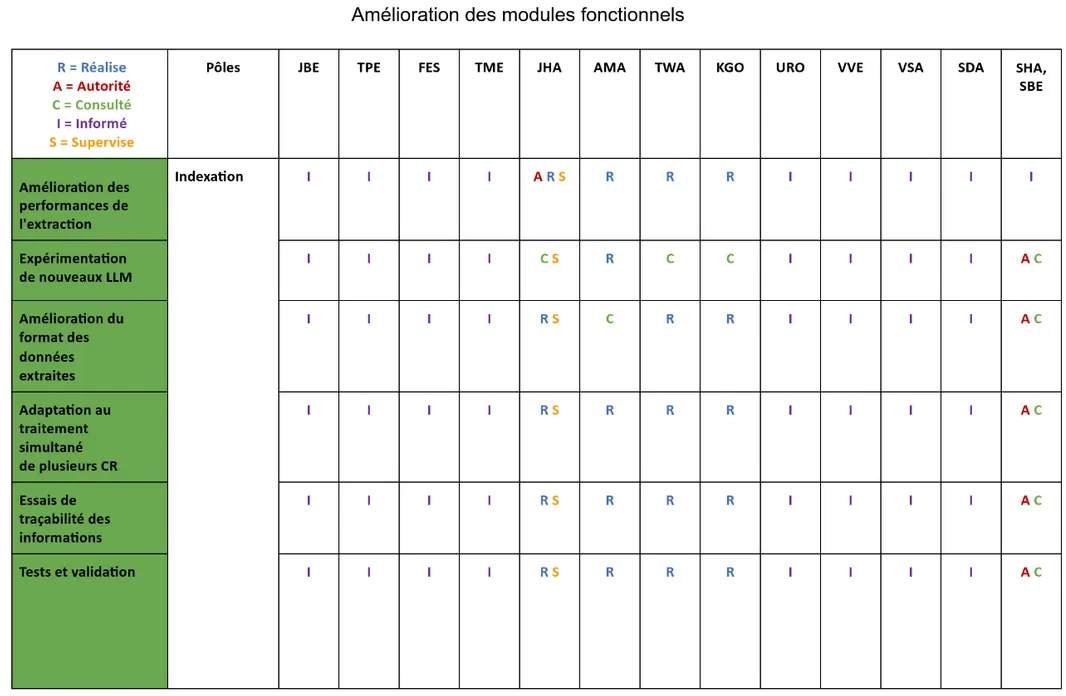
Figure 6 - RACI Matrix Excerpt
f) Training and Consultations
Throughout this project, we had the opportunity to work with numerous experts in their fields to help us progress. We therefore recorded all those we called upon and noted the total hours of training and consultation we used. In the end, we had used 14 hours of internal consultation (with people employed by the Centrale Lille group) and 23 hours of external consultation (with SIB representatives and medical staff).
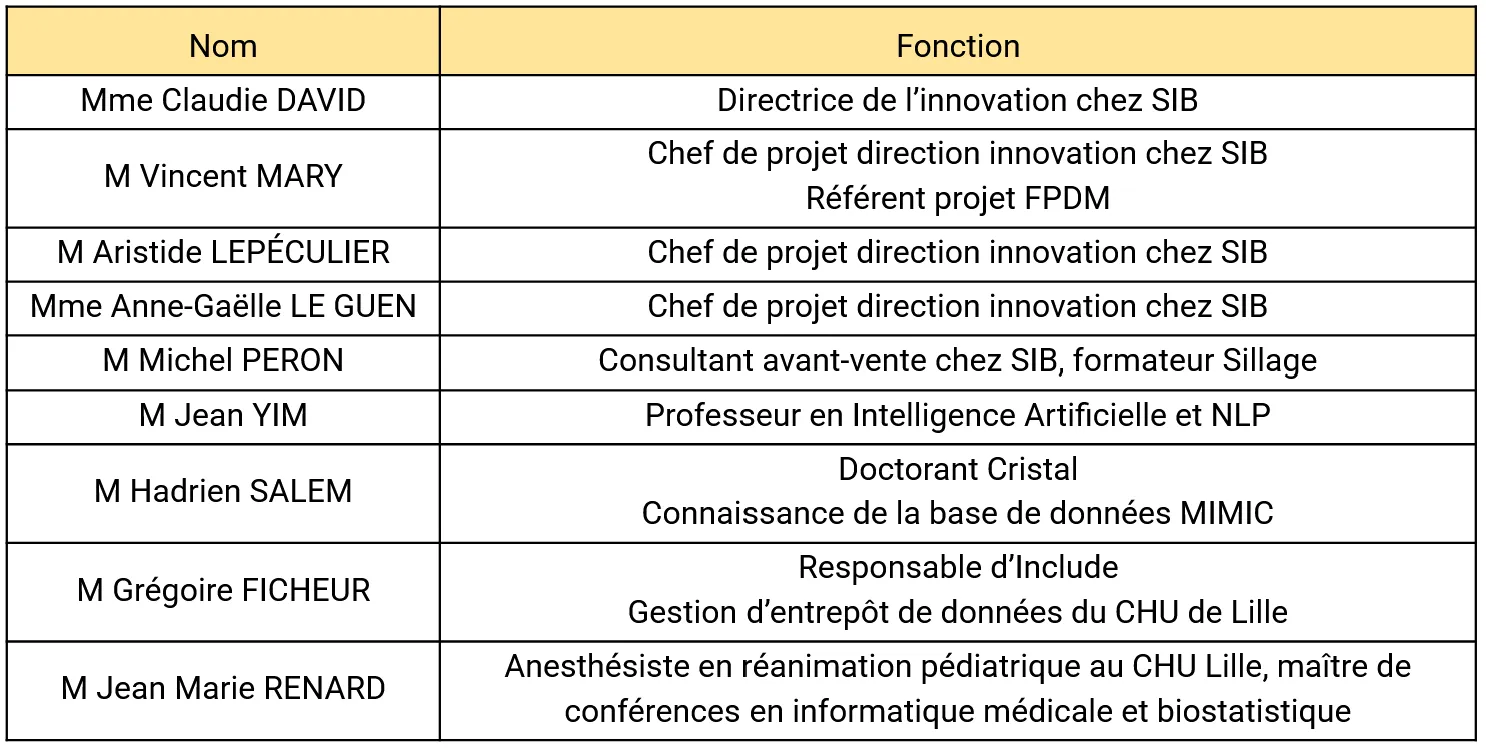
Figure 7 - All Consultants Met
g) Others
We mainly used 5 tools to organize our work:
- Discord: Communication
- Google Drive: Document sharing, collaborative work
- Hugging Face: Host for LLM and ML models used in different units
- Mimic IV: Medical database present on Cristal server
- GitHub: Collaborative environment for developing and managing versions
Finally, for our project to have a clear status, we drafted and signed an internal agreement with the Cristal laboratory.
3. Technical Deliverables
Due to the confidentiality surrounding the technical part of this project, I unfortunately cannot share the code we wrote. I will therefore be concise, but the implemented solution is much more complex than what I describe here.
a) Indexing Unit
The foundations of this work were built by synthesizing patient data into a text file from numerous joins on the MIMIC IV data table. These joins allowed us to keep only the most important parameters.
LLM Model Used: OpenHermes 2.5 - Mistral 7B
Objectives:
- Retrieval and concatenation of the entire patient medical record
- Synthesis and indexing of this data
- Work on result explainability
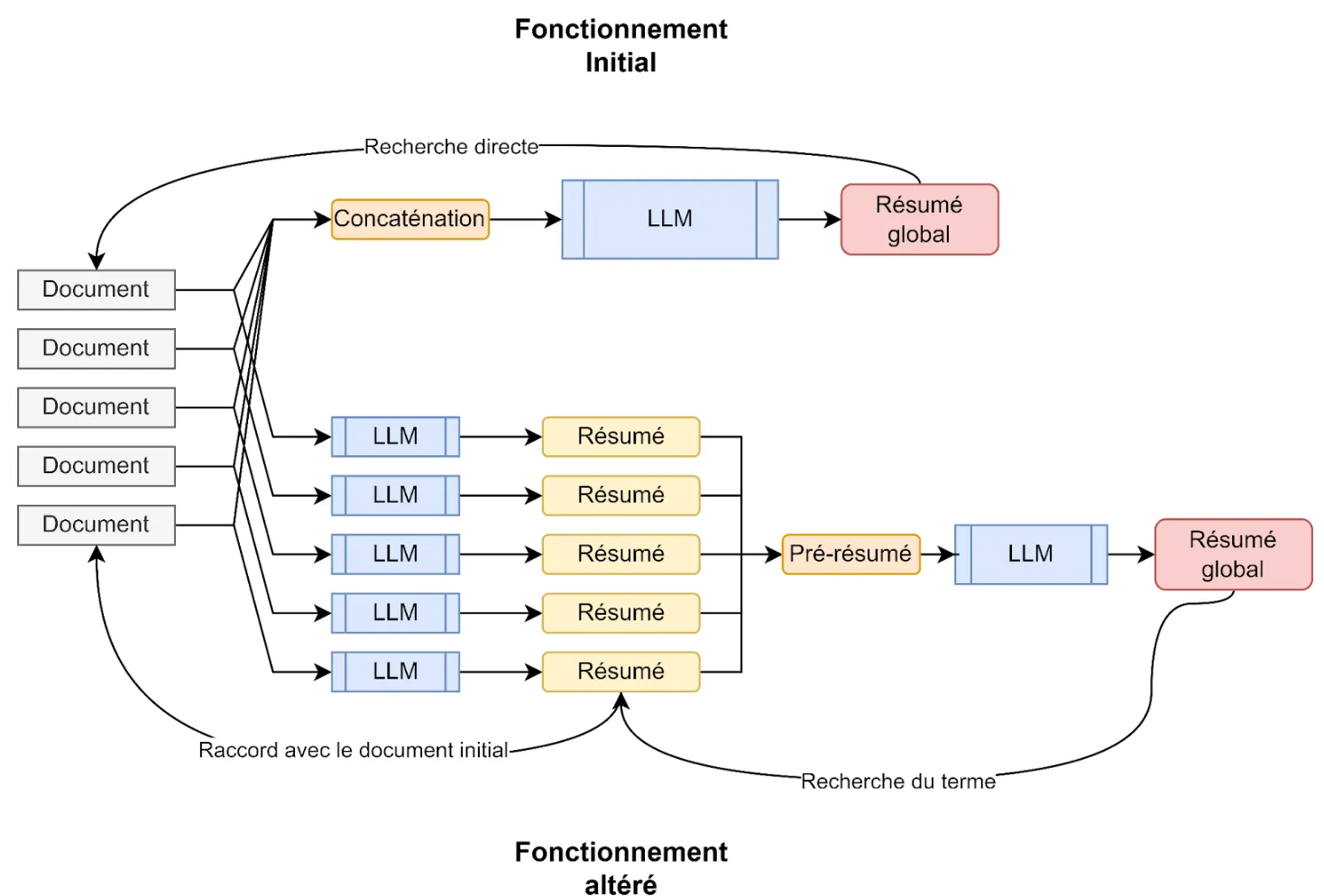
Figure 8 - Schematic of Our Solution’s Operation
Result: We finally manage to index patient data into a structured file

Figure 9 - Example of Report Before and After Indexing
b) Patient Triage Unit
To decide the triage result, several data points deemed to have a strong influence on patient priority level are retrieved:
| Temperature | float |
|---|---|
| HeartRate | int |
| RespiratoryRate | int |
| OxygenSaturation | float |
| BloodPressure | string |
| TransportMode | string |
| Age | int |
| Gender | string |
Models Used: OpenHermes 2.5 - Mistral 7B & XGBoostClassifier
Method:
- Structured decision tree to direct patients to appropriate care.
- Vectorization (Machine Learning), cosine similarity calculation
- LLM Prompt
- Python dictionary
c) Length of Stay Prediction Unit
Objectives:
- Predict length of stay at the time of admission to a department
- Useful for relatives and bed/staff management
- Based on reports written since hospital entry
Modules:
- Important word extraction with an LLM
- Prediction from examples to identify trends through information
- Training dataset containing examples of data with correct answers
Techniques Used:
- One-Hot Encoding for categorical variables
- StandardScaler for data normalization
- Principal Component Analysis (PCA) for dimensionality reduction
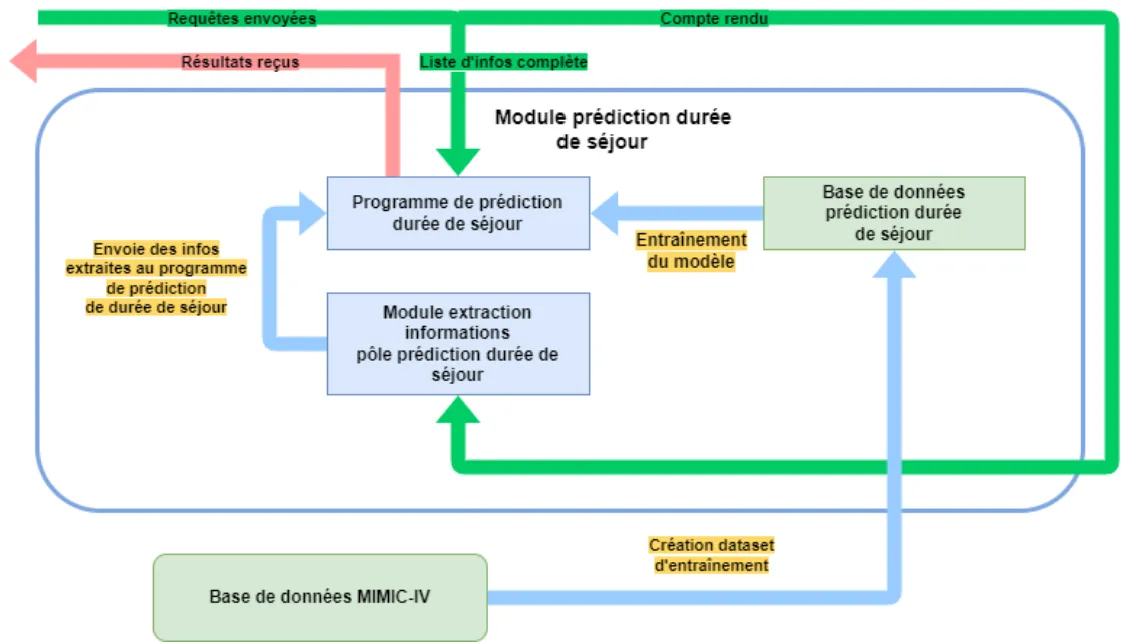
Figure 10 - Summary of Prediction Deliverable Operation
d) API Unit
Technologies Used: Flask, FastAPI, Django
Objectives:
- Allow hospital services to access predictions via an intuitive interface
- Manage access and permissions via a secure database
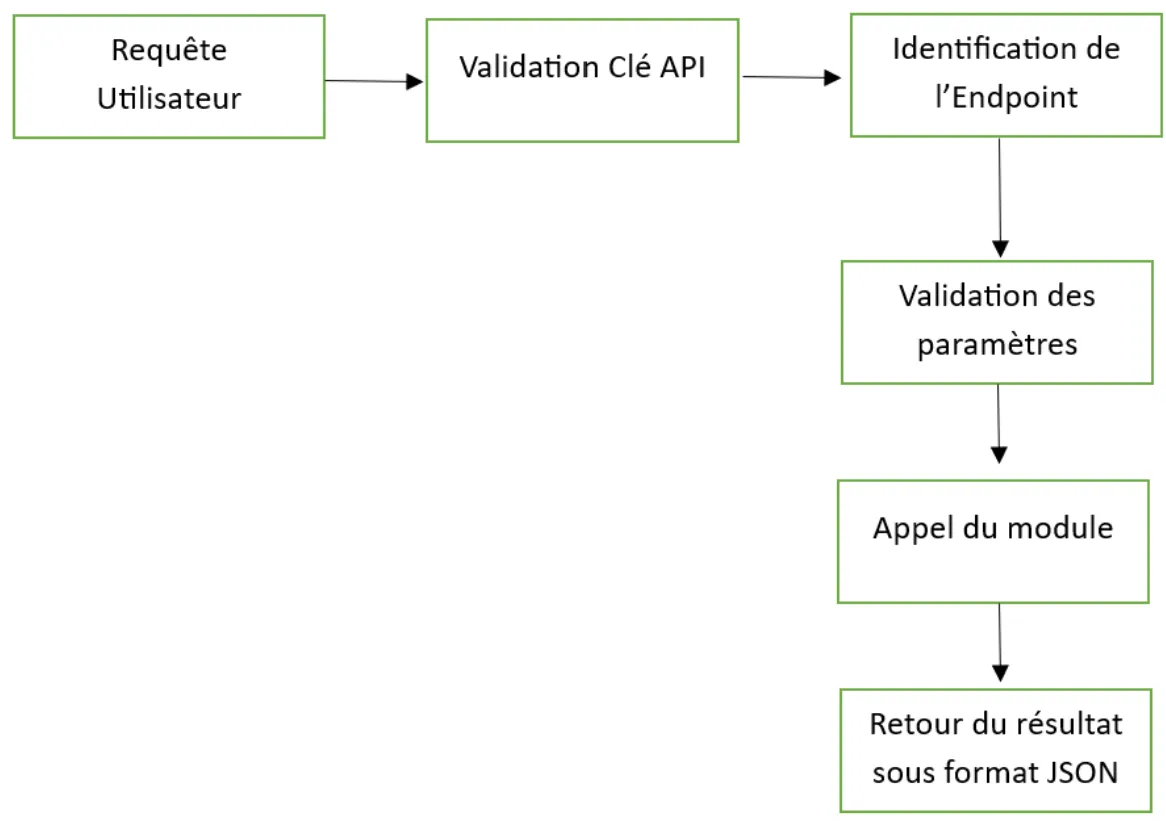
Figure 11: Different Steps of API Operation
4. Documentation and Final Deliverables
- Documented source code and API: Available on GitHub
- User and technical documentation: Detailed explanation of modules and usage guide
- Video demonstrations: Presentation of different implemented features
- Scientific article: In progress
These projects might interest you
Exploring a RAG Pipeline
RAG pipeline benchmark and assembly of the tool into a Chainlit chat.
MCP Data Science
102 data science tools driven by natural language via Anthropic's MCP protocol.